Optimization on Indexed Face Sets
The ifs subpackage of optimize contains
utilities to wrap functionals defined on
IndexedFaceSet
to work with the usual optimizers, and quick access to apply the minimizers
scipy.optimize.least_squares
and
scipy.optimize.minimize
of scipy.
Examples for functionals can be found in functionals.
Indexed Face Set functionals
IndexedFaceSet
are assumed to be functions of the formfunctional(indexedfaceset, attribute_name, *additional_attribute_names), e.g.def dirichlet(indexedfaceset, attribute_name):
e = np.int32(indexedfaceset.edges)
attribute_dict = indexedfaceset.vertex_attributes
attr_values = attribute_dict[attribute_name]
return attr_values[e[:, 0]] - attr_values[e[:, 1]]
Note that this function would return a #edges long vector. Since for some problems it might be useful to apply different norms, we expand our notion of functionals to include vector valued functions as well.
Preparing a functional for optimization
In order to work with most optimizers, functionals as defined above have to be
wrapped to accept a single vector as an input. For this we provide the wrappers
flatten_functional(),
flatten_functional_ignore(),
flatten_gradient(), and
flatten_gradient_ignore(),
that reshape and store this vector in an attribute of an indexed face set before
evaluating the functional/gradient.
import ddg
import numpy as np
faces = [[0,1,4,3],
[1,2,5,4],
[3,4,7,6],
[4,5,8,7]]
u = np.array([0, 0, 0, 0, 1, 0, 0, 0, 0], dtype=float)
ifs = ddg.datastructures.indexedfaceset.ifs.IndexedFaceSet(faces)
ifs.set_attribute('u', 'verts', np.copy(u))
wrapped_dirichlet = ddg.optimize.ifs.flatten_functional(ifs, dirichlet, 'verts', 'u',
boundary=[0, 1, 2, 3, 5, 6, 7, 8])
result = dirichlet(ifs, 'u')
assert (result @ result == 4.)
Here we also marked all vertices except 4 as boundary cells.
By using flatten_functional() the wrapped function
requires a single value. If we were to use
flatten_functional_ignore instead, we would be required to pass a value
for each vertex, but all except the value for vertex 4 would be ignored.
ign_dirichlet = ddg.optimize.ifs.flatten_functional_ignore(ifs, dirichlet, 'verts', 'u',
boundary=[0, 1, 2, 3, 5, 6, 7, 8])
result = ign_dirichlet(u)
Note that calling the wrapped function changes the attribute values in place
ifs.vertex_attributes['u'][4] == 1.
result = wrapped_dirichlet(0)
assert (result @ result == 0.)
for val in ifs.vertex_attributes['u']:
assert (val == 0.)
wrapped_dirichlet(1.)
array([ 0., 0., -1., 0., 0., -1., 0., 1., 0., 0., 1., 0.])
Optimizing a functional with scipy
Now that our functional handles a single vector as its input, we can use external optimizers for it.
import scipy.optimize
optimize_result = scipy.optimize.least_squares(wrapped_dirichlet, (4,))
optimize_result.x
array([0.])
Using this method to optimize a functional on an indexed face set, the current values stored on it will generally not be the actual result of the optimization, since optimizers like those provided by scipy will approximate the gradient of our functional at this point to check for their break condition (usually |gradient(p)| < tol). This is done by evaluating the functional at a couple of points near the current result, which causes additional writes to the attribute array of our indexed face set. To ensure that the value is written on the indexed face set, we can reevaluate the wrapped functional with the optimization result.
assert (not ifs.vertex_attributes['u'][4] == 0.)
result = wrapped_dirichlet(optimize_result.x)
assert (result @ result == 0.)
assert (ifs.vertex_attributes['u'][4] == 0.)
Another way to use scipy’s optimization is given by the functions
minimize() and least_squares() which use
the respective optimizers of the same name provided by scipy. They also ensure that the result
of the optimization is written back to the indexed face set attribute.
ifs.set_attribute('u', 'verts', np.copy(u))
optimize_result2 = ddg.optimize.ifs.least_squares(dirichlet, 'verts', 'u'
boundary=[0, 1, 2, 3, 5, 6, 7, 8])
optimize_result2.x
array([0.])
assert (ifs.vertex_attributes['u'][4] == 0.)

Fig 1. Graph of attribute u before… |

Fig 2. …and after optimization |
Examples
Optimizing with a gradient
Giving an optimizer a gradient generally ensures that it does not approximate it instead, and thus in most cases the result of the optimization is already present on the indexed face set after the optimization process is completed.
from ddg.optimize.ifs.functionals import flat_cubic_quad, g_flat_cubic_quad
import ddg.optimize.ifs as optf
co = np.array([[0,0,0],
[0,1,0],
[0,2,0],
[1,0,0],
[1,1,1],
[1,2,0],
[2,0,0],
[2,1,0],
[2,2,0]], dtype=float)
ifs.set_attribute('co', 'verts', np.copy(co))
wrapped_functional = optf.flatten_functional(ifs, optf.functionals.flat_cubic_quad,
'verts', 'co', boundary=[0,1,2,3,5,6,7,8])
wrapped_gradient = optf.flatten_gradient(ifs, optf.functionals.g_flat_cubic_quad,
'verts', 'co', boundary=[0,1,2,3,5,6,7,8])
optimization_result = scipy.optimize.least_squares(wrapped_functional, (1., 1., 1.),
jac=wrapped_gradient)
assert (np.isclose(ifs.vertex_attributes['co'][4], optimization_result.x).all())
result = flat_cubic_quad(ifs, 'co')
assert (result @ result < 1e-15)
Flattening a bell
We start of by creating a 50x50 grid and sample the density \(\mathit{u}\) of a normal distribution on \([-1, 1]^2\).
def density(x):
return 2 * (1/np.sqrt(2*np.pi) * np.e**(-x**2*10))
faces = []
for row in range(49):
for col in range(49):
base = col + row*50
faces.append([base, base+1, base+n+1, base+n])
grid = ddg.datastructures.indexedfaceset.ifs(faces)
coords = np.linspace(-1, 1, 50)
gridco = np.array([[coords[divmod(i, 50)[1]], coords[divmod(i,50)[0]] for i in range(2500)])
u = np.prod(density(gridco), axis=1)
bellco = np.column_stack([gridco, u])
grid.set_attribute('u', 'verts', u)
grid.set_attribute('bellco', 'verts', bellco)
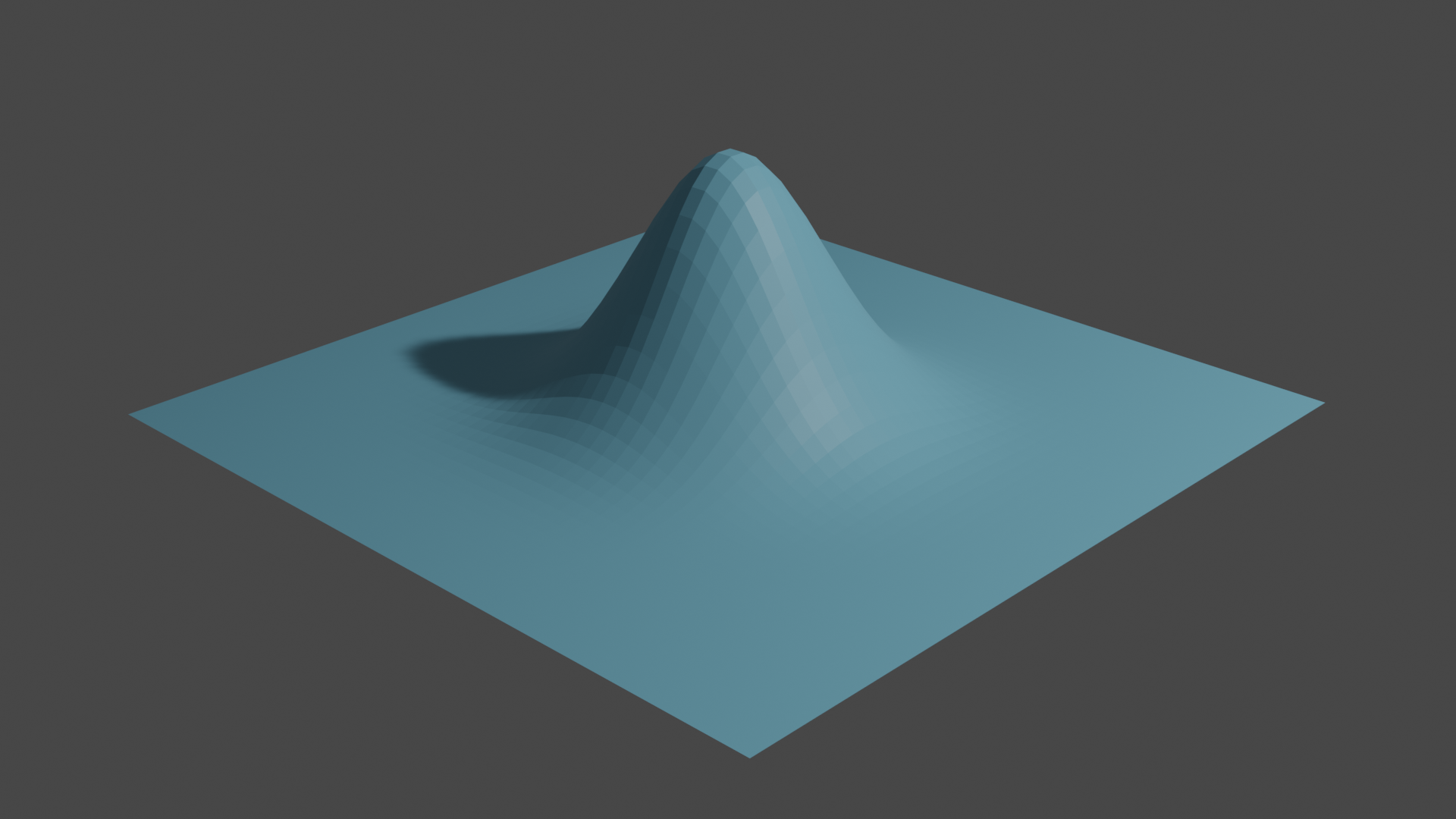
Fig 3. The graph of \(\mathit{u}\) over our grid
Next we minimize the Dirichlet energy of \(\mathit{u}\) while fixing \(\mathit{u}\) along the boundary of the grid.
boundary1 = np.arange(50)
boundary2 = boundary1 + 50 * 49
boundary3 = np.arange(50, 50 * 49, 50)
boundary4 = np.arange(2 * 49, 50 * 49, 50)
boundary = np.append(boundary1, boundary2)
boundary = np.append(boundary, boundary3)
boundary = np.append(boundary, boundary4)
wrap_dirichlet = optf.flatten_functional(grid, optf.functionals.dirichlet,
'verts', 'u', boundary=boundary)
initial = optf.utils.initial_guess(grid, 'verts', 'u', boundary=boundary)
res = scipy.optimize.least_squares(wrap_dirichlet, initial)
wrap_dirichlet(res.x) # re-evaluating writes the correct result back on the grid
grid.set_attribute('opt', 'verts', np.column_stack([gridco, grid.vertex_attributes['u']))
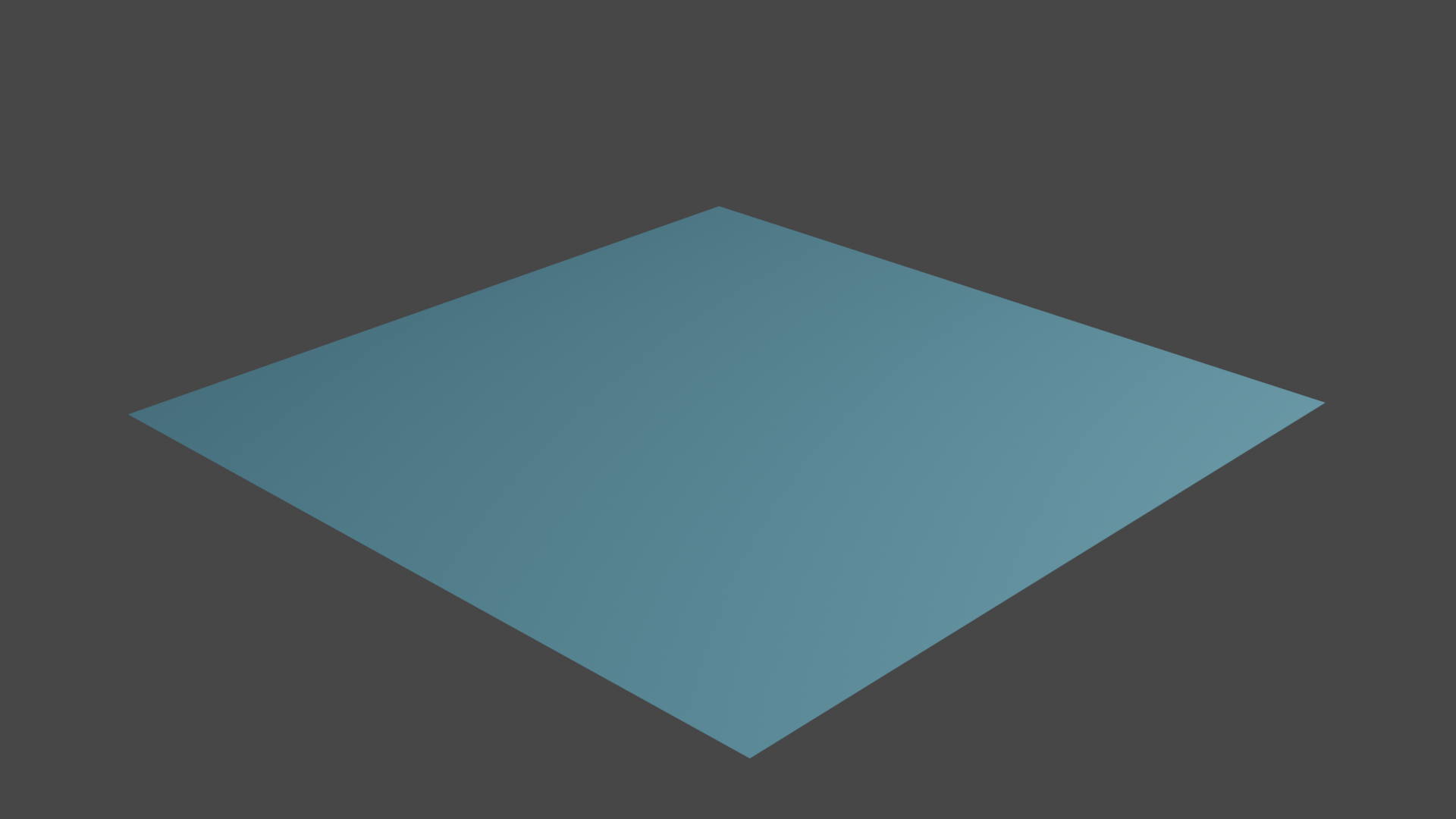

Fig 4. Optimization result